Building a Protein-Understanding LLM: A Week of Trials, Errors, and Breakthroughs
The story of building a multimodal LLM that processes protein embeddings alongside natural language—from architecture decisions to debugging disasters to assembling 4.5 million training records.
What happens when you bridge protein language models with large language models? Can we make an LLM that leverages structural protein embeddings, not just parses their sequences?
The Beginning: Why This Project?
It started with a simple observation. Protein language models like ESM-3 have learned remarkably rich representations from millions of protein sequences—structural features, evolutionary constraints, functional signals—all encoded in high-dimensional embeddings. Meanwhile, LLMs excel at reasoning, following instructions, and generating coherent explanations. But they exist in separate worlds.
The question that kept nagging me: what if we could bridge these two worlds?
Not by training a protein model from scratch (prohibitively expensive), but by post-training—freezing the protein encoder, attaching a learnable bridge, and teaching an LLM to interpret protein embeddings alongside natural language. The best of both worlds, without the compute bill.
This post documents the journey—the architectural choices, the debugging disasters, the data struggles, and what we learned along the way.
Day 1: Architecture Decisions
The Core Design
We settled on a modular pipeline:
- Protein Encoder (frozen): ESM-3 small with 1.4B parameters producing 1536-dimensional embeddings
- Attention Pooling: Compress variable-length residue embeddings into 32 fixed tokens
- MLP Projector (trainable): Map 1536-dim protein space into 2560-dim LLM input space
- LLM: Qwen3-4B with LoRA on all linear layers (attention and MLP)
The idea is elegant: ESM-3 handles the protein understanding, the projector translates between representation spaces, and the LLM handles reasoning and generation. We keep ESM-3 frozen to preserve its hard-won protein knowledge, and use LoRA to efficiently adapt the LLM.
Start with 32 pooled tokens. Too few and we lose protein information; too many and we blow up sequence length. 32 seemed like a reasonable middle ground—equivalent to roughly 100 words of text context.
The Three Encoding Approaches
We actually designed three approaches to compare:
Raw sequence as <protein>MKTL...</protein> — the simplest baseline. No ESM-3, just treating proteins as weird text.
ESM-3 embeddings → Attention Pooling → MLP Projector. Our main approach.
ESM-3 embeddings → Perceiver Resampler. ~29M parameters, comparable to MLP, with latent_dim decoupling for potentially better information compression.
We later added a fourth approach — Flamingo-style gated cross-attention — bringing the comparison to four pathways.
The four-way comparison would be our core thesis experiment. But first, we had to make anything work at all.
Day 2: First Training Run — It Works!
The Mol-Instructions Dataset
We started with Mol-Instructions, a curated set of 505K protein instruction pairs covering five task types:
| Task | Samples | Description |
|---|---|---|
| Catalytic Activity | 53K | Predict enzyme reactions |
| Protein Function | 114K | Predict biological roles |
| General Description | 87K | Describe protein properties |
| Domain/Motif | 45K | Identify structural domains |
| Protein Design | 196K | Generate sequences (output, not input) |
The format is clean: {instruction, input, output} where the input contains the protein sequence. Perfect for supervised fine-tuning.
First Results
| Run | Samples | Loss (Start → End) | Duration |
|---|---|---|---|
| 500-sample test | 500 | 17.35 → 4.08 | ~6 min |
| 10K baseline | 10,000 | 35.80 → 27.84 | ~96 min |
| 50K full | 50,000 | 34.25 → 14.77 | ~2.35 hr |
The training converged. On 8x H100 80GB GPUs, the 50K run used only ~39 GB memory with an effective batch size of 32. The architecture was functional.
But then came the evaluation…
Day 2-3: The Evaluation Reality Check
What Works
The vanilla Qwen3-4B (before any fine-tuning) set a floor. It had never seen proteins explicitly, so performance was essentially random:
- GO Term Prediction: F1 Micro = 0.047 (the model over-predicted 11.9 terms vs 5.0 ground truth)
- PPI Prediction: AUROC = 0.51 (literally random)
- Stability Prediction: Pearson r = 0.094 (no correlation)
After SFT, the numbers… didn’t improve much. In fact, some got worse.
GO prediction degraded after SFT. The vanilla model at least guessed some correct terms among its many predictions. After SFT, the model predicted too few terms (or none at all). F1 went from 0.047 to 0.0.
The PPI Bias Problem
All models, regardless of scale (4B, 8B, 14B), defaulted to predicting “No interaction” for every protein pair. High specificity (0.9-1.0), terrible recall (0.2 or 0.0). The models were stuck at majority-class prediction.
The silver lining: AUROC improved with scale (0.51 → 0.60 → 0.70), suggesting the models learned better internal representations even when failing to express positive predictions.
What This Told Us
- The architecture works — training converges, memory is manageable
- But SFT alone isn’t enough — instruction-tuning doesn’t automatically teach structured output (GO terms) or overcome class imbalance (PPI)
- Scale helps internal representations — the knowledge is there, it just doesn’t surface in generation
We needed two things: better training data and task-specific rewards (RL).
Day 4: The Instruct Model Disaster
This is the debugging war story.
We launched an 8B training run overnight. The next morning, the loss curve looked fine:
| Step | Train Loss | Eval Loss |
|---|---|---|
| 250 | 8.30 | 2.09 |
| 1000 | 4.80 | 1.89 |
| 5000 | 3.86 | 1.84 |
But generation was garbage. Literal garbage:
KQKQKQKQKQKQKQKQ...
SSSSSSSSSSSSSSSS...
(empty string)
After hours of debugging, I found it: the config file had path: Qwen/Qwen3-8B — the base model, not Qwen/Qwen3-8B-Instruct-2507.
Base models don't know how to stop. They never learned instruction-following, chat templates, or turn-taking. The LoRA adapter we trained was bound to the base model's weight space and couldn't be transferred to Instruct. The entire overnight run was wasted.
The Fix
All model configs now use -it suffix to make this mistake impossible:
| Config | Before | After |
|---|---|---|
| qwen3_4b.yaml | qwen3_4b | qwen3_4b_it |
| qwen3_8b.yaml | qwen3_8b | qwen3_8b_it |
| qwen3_14b.yaml | qwen3_14b | qwen3_14b_it |
A simple naming convention that would have saved 8 hours of H100 time. Lesson learned.
Day 4-5: Protein Boundary Tokens
The Problem
The ESM-3 embedding path used a single <|protein_embed|> placeholder. The LLM had no explicit signal for where the protein representation starts and ends. It was just… 32 tokens appearing in the middle of text.
The Solution
We introduced structured boundary tokens:
<|protein_start|> <|protein_embed|> <|protein_end|>
ID 151669 ID 151670 ID 151671
At forward time, the embedding looks like:
[..., text, START_embed, prot_1, prot_2, ..., prot_32, END_embed, text, ...]
↑ LLM learns ↑ 32 ESM-3 pooled tokens ↑ LLM learns
what this means (replaced at forward) what this means
The start/end tokens remain as regular LLM embeddings—the model learns what they mean through training. The middle tokens get replaced by ESM-3’s pooled output. All three are masked from loss computation.
This gave the LLM clear modality boundaries, and the next training run showed noticeably better attention patterns on the protein regions.
Day 6: Scaling Up Data
Why 505K Wasn’t Enough
Mol-Instructions covers five tasks well, but they all come from one paper’s annotation pipeline. The model learned that particular group’s phrasing from that particular subset of UniProt. We needed:
- Different annotation styles: Long-form paragraphs vs. structured one-liners
- More tasks: Subunit structure, tissue specificity, PTMs, gene prediction
- Scale: From ~300K effective records to millions
The Six Sources
After evaluating several candidates (dropping two for noise/unavailability), we assembled:
| Source | Records | What It Brings |
|---|---|---|
| Mol-Instructions | 299K | Curated instruction pairs (ICLR 2024) |
| Swiss-Prot | 1.08M | Gene prediction, organism prediction, function |
| ProteinLMDataset | 826K | Subunit, PTM, disease, tissue, induction |
| SwissProtCLAP | 511K | Rich paragraph-length descriptions |
| ProtDescribe | 1.76M | Naming, similarity, location, function |
| Protein2Text-QA | 52K | 44,915 unique questions — highest diversity |
Total: 4.52 million training records.
Sequence Length Distribution
One thing to always check with protein datasets: the length distribution. Protein length controls ESM-3 memory (geometric attention is O(L²)), training sequence length, and representation quality.
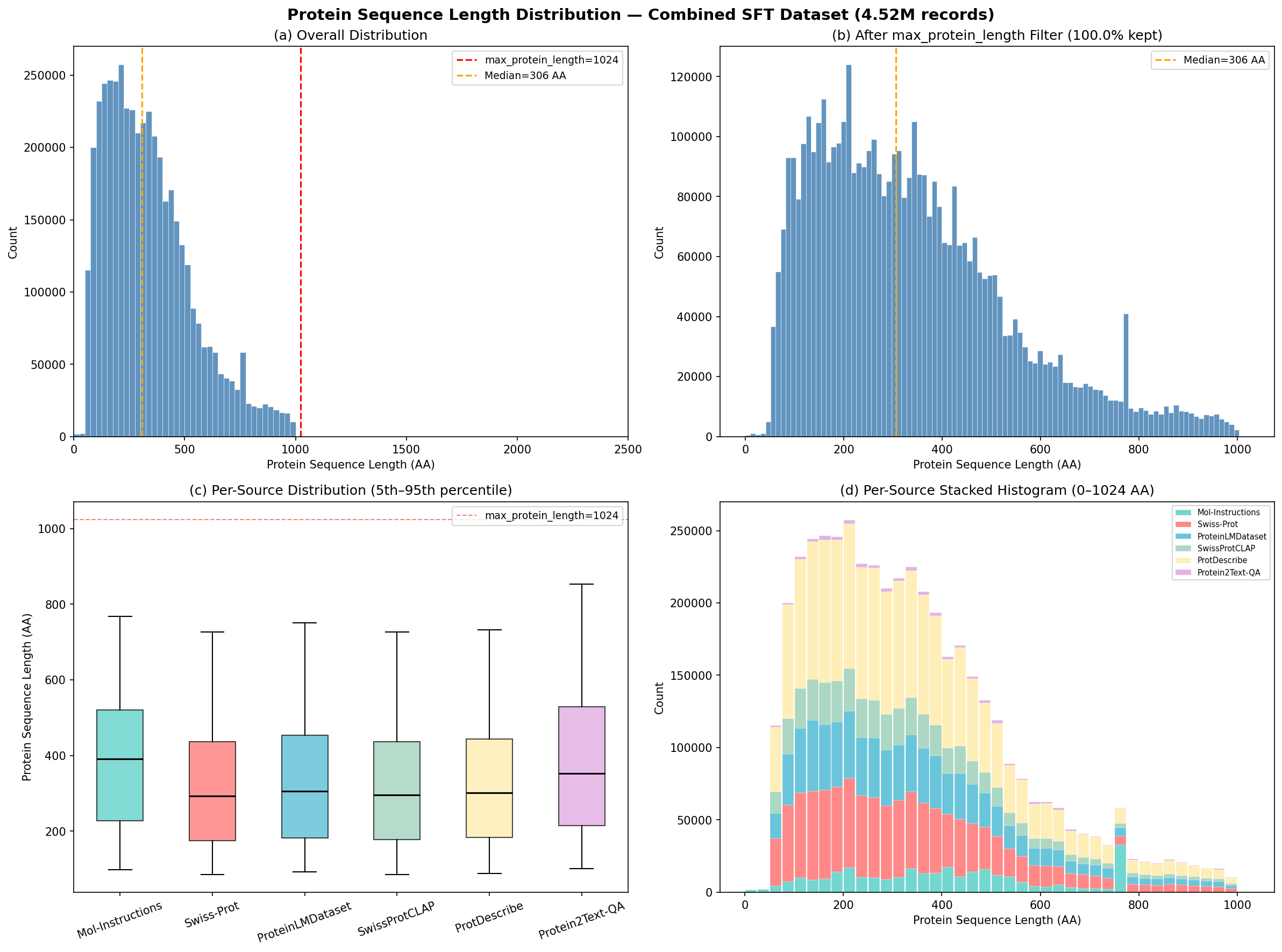
Distribution of protein sequence lengths across the combined 4.5M dataset. The median sits at 306 amino acids.
| Statistic | Value |
|---|---|
| Median | 306 AA |
| Mean | 339 AA |
| 5th percentile | 89 AA |
| 95th percentile | 751 AA |
| Max | 1,000 AA |
The distribution peaks at 150-350 amino acids (typical single-domain proteins) with a long right tail. Mol-Instructions had a hard cutoff at 768 AA; the other sources extend to 1,000 AA—introducing longer proteins the model hadn’t seen before.
Data Quality Gotchas
Single-word outputs: 31% of ProteinLMDataset subunit records have outputs like just "Monomer." — accurate, but not the verbose responses we want. 12-16% duplicates: Template augmentation inflated effective weight of some sources. 72.8% sequence overlap: Most sources derive from UniProt, so the same ~440K proteins appear repeatedly—but in different analytical contexts.
The RL Path Forward
SFT established the foundation, but the evaluation results made clear: we need task-specific rewards to push beyond “reasonable-sounding but wrong” predictions. We implemented GRPO (Group Relative Policy Optimization) with four verifiable reward functions — GO term F1, stability prediction accuracy, PPI interaction detection, and ESMFold structural quality assessment. No reward model needed, just computable metrics from model outputs.
The full GRPO story — including a surprising reversal where text-only outperforms ESM-3 — is told in Part 3.
Current State and What’s Next
Where This Leads
The architecture works, the data pipeline works, and the evaluation framework is in place. The next chapters of this story:
- Part 2: Scaling to 4.9 Million Samples — What happens when we go from 50K to millions of training examples across six data sources
- Part 3: The Gradient Routing Problem — GRPO reinforcement learning reveals a fundamental tension between the encoder bridge and LLM adapters
- Series Overview — The full project arc, including what’s next
Resources
Project Repository: Post_Training_Protein_LLM
Related Work:
Related Posts:
